The TRUPCR® NRAS Mutation Kit is an in vitro diagnostic test intended for the qualitative detection of NRAS somatic mutations in the genomic DNA extracted from fresh, frozen or formalin fixed paraffin-embedded (FFPE) tissue. The TRUPCR® NRAS mutation kit is based on allele specific amplification and is achieved by ARMS PCR. Taq DNA polymerase is extremely effective at distinguishing between a match and a mismatch at the 3’-end of a PCR primer. The kit is designed to selectively amplify mutant specific sequences in samples that contain a mixture of wild-type and mutated DNA. The most common mutations are found in codon 12, 13 and 61. The detection is achieved using fluorescent probes labelled with FAM and HEX. The TRUPCR® NRAS Kit is composed of 11 assays for the detection of the NRAS mutations and a reference control gene of NRAS region without any known polymorphism/mutation.
Key Features:
- Selective Amplification of DNA containing mutation with ARMS Technology.
- Sensitive to detect up to 1% mutation in NRAS gene.
- Detects 24 different mutations in a single run.
- Extraction control included to avoid false-negative results.
- Rapid, more reliable, comprehensive and cost effective tests.
- Easy work flow & compatible with various Real Time PCR instruments.
NRAS Detectable mutations:
| TUBE | CODON | MUTATION | NUCLEOTIDE CHANGE | REMARKS |
|---|---|---|---|---|
| G12ACD PPM (1) | 12 | G12A | c.35G>C | It detects 3 mutations but does not distinguish between them. |
| G12C | c.34G>T | |||
| G12D | c.35G>A | |||
| G12RSV PPM (2) | 12 | G12R | c.34G>C | It detects 3 mutations but does not distinguish between them. |
| G12S | c.34G>A | |||
| G12V | c.35G>T | |||
| G13ACD PPM (3) | 13 | G13A | c.38G>C | It detects 3 mutations but does not distinguish between them. |
| G13C | c.37G>T | |||
| G13D | c.38G>A | |||
| G13RSV PPM (4) | 13 | G13R | c.37G>C | It detects 3 mutations but does not distinguish between them. |
| G13S | c.37G>A | |||
| G13V | c.38G>T | |||
| A59X PPM (5) | 59 | A59T | c.175G>A | It detects 2 mutations but does not distinguish between them. |
| A59D | c.176C>A | |||
| Q61X PPM (6) | 61 | Q61H | c.183A>T | It detects 3 mutations but does not distinguish between them. |
| Q61H | c.183A>C | |||
| Q61P | c.182A>C | |||
| Q61K PPM (7) | 61 | Q61K | c.181C>A | |
| Q61L PPM (8) | 61 | Q61L | c.182A>T | |
| Q61R PPM (9) | 61 | Q61R | c.182A>G | |
| K117X PPM (10) | 117 | K117R | c.350A>G | It detects 3 mutations but does not distinguish between them. |
| K117N | c.351G>T | |||
| K117N | c.351G>C | |||
| A146T PPM (11) | 146 | A146T | c.436G>A | |
| NRAS Reference | Detects NRAS region without any polymorphism/mutation. | |||
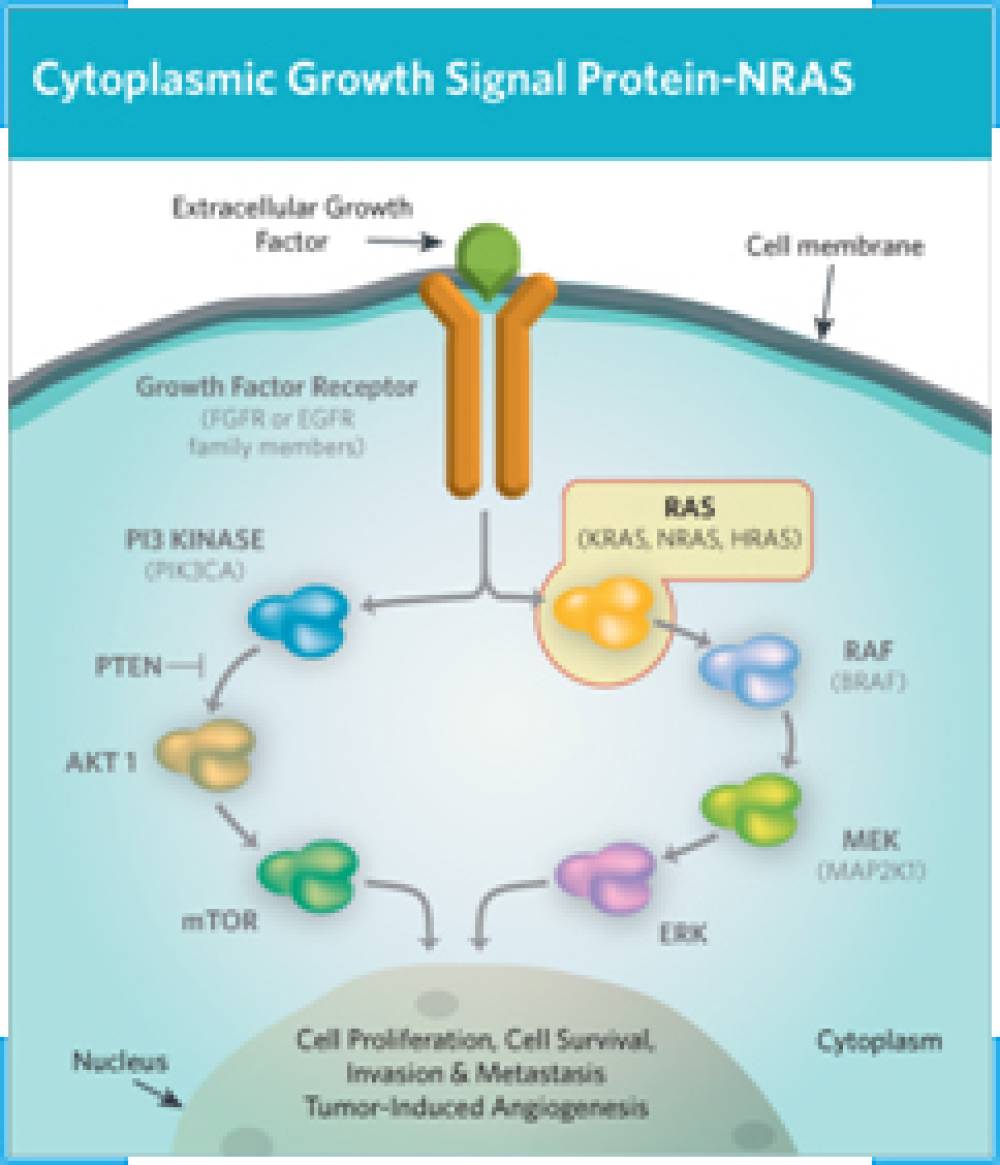
The RAS oncogenes (KRAS, NRAS, HRAS) encode a family of GTP-regulated switches that are recurrently mutated in human cancer. The four enzymes encoded by the three RAS genes are highly homologous to one another. Activating mutations render the RAS protein in the GTP-bound activated form by preventing hydrolysis of GTP and so preventing transition to the GDP-bound inactive state. RAS mutations occur at conserved hotspots, and oncogenic mutations have been shown to occur at codons 12, 13, 61, 117, and 146. Activation of the RAS proteins leads to pleiotropic effects in cells, including cellular proliferation, survival, and differentiation. The differences in signaling and downstream effectors that result from the different activated RAS isoforms are not currently clear. In colorectal cancer (CRC), RAS mutations predominantly occur in the KRAS gene; 45% of metastatic CRC (mCRC) harbor an activating KRAS mutation. NRAS mutations occur in 2-7% of mCRC cases.
NRAS mutations are now regularly identified in the clinical treatment of mCRC as part of routine RAS testing prior to EGFR (epidermal growth factor receptor) inhibitor therapy. The clinical implications of NRAS mutation beyond lack of response to anti-EGFR therapy (4) and whether these tumors behave similarly to KRAS mutant mCRC are not known. KRAS mutations have been associated with right-sided colon tumors, while NRAS mutations have been associated with left-sided primary tumors and female gender(3), suggesting a distinct biology for KRAS and NRAS mutant molecular subsets of mCRC.
Ordering Information
| CAT. NO. | PRODUCT | CONTENTS |
|---|---|---|
| 3B1273 | TRUPCR® NRAS Mutation Kit | 48 Reactions |
| 3B1274 | TRUPCR® NRAS Mutation Kit | 96 Reactions |